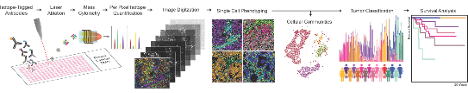
To understand tissues and tumours as the integrated outcome of their single cell components, we utilize multiplexed imaging to simultaneously quantify single cell phenotypes and markers of their functional state, as well as their interactions, overall organization, and contribution to tissue architecture. These methods facilitate spatially-resolved screening of distinct clones in mouse models of disease and the identification of biomarkers associated with clinical outcome in biobanked patient tissues.
Understanding how gene/protein/cell networks contribute to health and disease requires an understanding of the spatially resolved quantification and modeling of many biomarkers at different scales, from subcellular to their overall expression in single cells and organized tissues. For this purpose we use a combination of quantitative, spatially-resolved multiplexed technologies with a focus on Imaging Mass Cytometry (IMC).
Projects
Precision medicine
In clinical patient samples, we study the multi-cellular contents of healthy tissues and diverse tumour types and study the cellular phenotypic changes that take place during disease progression and in response to therapy. We focus on the retrospective analysis of large biobanked cohorts or the prospective study of small biopsy tissue samples, both of which present unique technical challenges. We aim to identify cell states and multi-cellular networks that can inform clinical decision making and be targeted for therapeutic intervention.

Pancreatic Cancer: As part of the Pancreatic Cancer Translational Research Initiative at the Ontario Institute of Cancer Research, we are studying how intra- and inter-tumour cellular heterogeneity contributes to therapeutic resistance and poor outcomes.
https://oicr.on.ca/programs/pancreatic-cancer-translational-research-initiative-pancurx/
Breast Cancer: By pairing mouse models of mammary tumorigenesis and highly multiplexed cellular profiling of human breast cancers we are investigating the molecular pathways which drive breast cancer initiation and metastasis. For this purpose we are working closely with scientific collaborators within the Lunenfeld Tanenbaum Researcher Institute and the clinical teams who have created the Breast Translational Research Resource at Mount Sinai Hospital.
Kidney Cancer: Through our US DOD funded partnership with the urology surgical team at Toronto General Hospital we are investigating the immune cell networks that regulate clear cell renal cell carcinoma (ccRCC) and their relationship to immunotherapy response and resistance.
Cancer initiation and metastasis Utilizing a combination of genetic tools and spatially-resolved quantitative single cell approaches, in vivo we are tracking the development of cancer from the first cells which initiate cancer to localized sites of dissemination and the distant metastases they form. In doing so, we are working to identify and target the molecular bottlenecks that mitigate advanced disease.
Tumour regulation of its microenvironment In both human disease and in controlled model systems, we are investigating how tumours regulate their local microenvironment and influence immune cell recruitment. As part of the Terry Fox Research Institute Hippo Pathway Team we have developed a spatially resolved functional genomics screening approach to dissect the extracellular signaling networks controlled by tumour-specific Hippo signaling.
http://Hippo pathway PPG team: https://www.tfri.ca/our-research/research-project/targeting-the-hippo-signaling-network-in-cancer
Our work: https://www.tfri.ca/our-research/research-project/characterization-and-treatment-response-of-hippo-pathway-regulated-tumour-microenvironments
Single cell systems biology methods development
We develop spatially resolved systems biology tools and analysis methods to quantify complex tissue systems across biological scales from the modification of a single gene, to genomes, transcriptomes and pathway specific signaling, and how these influence cellular phenotype and tissue level patterning. With specific expertise in imaging mass cytometry and CyTOF technology, we combine multiplexed imaging and spatially resolved ‘omics’ methods to investigate the molecular underpinnings of classic histopathology and unravel the complexity of tumour morphology and architecture. We are continually improving the quantitative methods we use, developing new reagents to measure cellular function, and creating new functional and phenotypic screening approaches. For example, as part of the Medicine by Design Grand Challenges program we are working with a multidisciplinary team of synthetic biologists, chemists, bioengineers, and computational biologists to engineer scalable multi-cellular environments.
https://mbd.utoronto.ca/research/grand-questions-program/
Funding
With thank all of our funding sources for their generous support of our work.